PyNTA: Python Nanoparticle Tracking Analysis¶
Nanoparticle tracking analysis refers to a technique used for characterizing small objects optically. The base principle is that by following the movement of nanoparticles over time, it is possible to calculate their diffusion properties and thus derive their size.
PyNTA aims at bridging the gap between experiments and results by combining data acquisition and analysis in one simple to use program.
PyNTA is shipped as a package that can be installed into a virtual environment with the use of pip. It can be both triggered with a built in function or can be included into larger projects.
Installing¶
The best place to look for the code of the program is the repository at https://github.com/nanoepics/pynta. In short, if you want to install PyNTA you can run the following command:
pip install git+https://github.com/nanoepics/pynta
If you need further assistance with the installation of the code, please check Installing.
Start the program¶
After installing, the program can be started from the command line by running the following:
python -m pynta -c config.yml
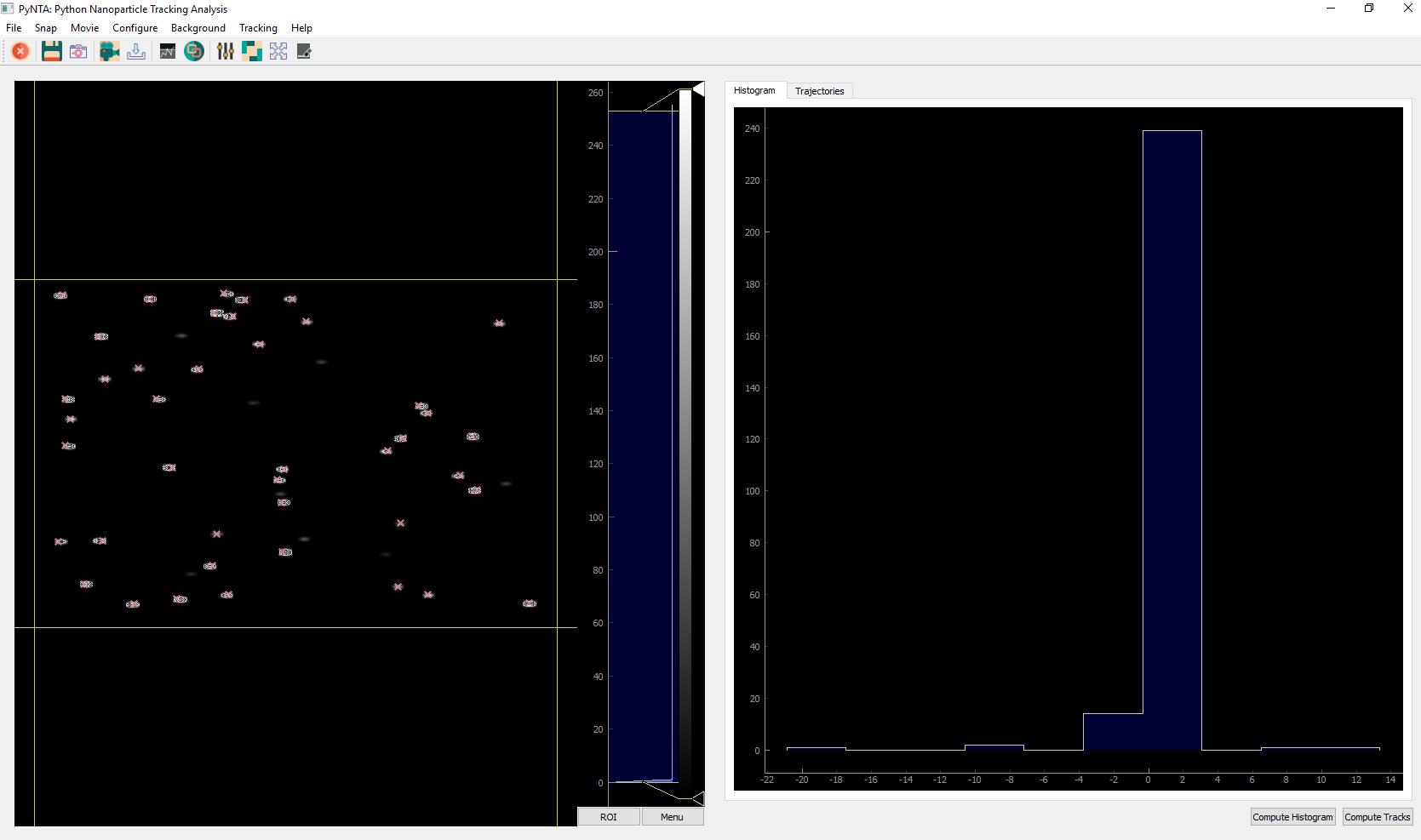
Remember that config.yml needs to exist. To create your own configuration file, you can start with the example provided in the examples folder. Once the program starts, it will look like the following:
Contributing to the Program¶
The program is open source and therefore you can modify it in any way that you see fit. You have to remember that the code was written with a specific experiment in mind and therefore it may not fulfill or the requirements of more advanced imaging software.
However the design of the program is such that would allow its expansion to meet future needs. In case you are wondering how the code can be improved you can start by reading How to Contribute Code, or directly submerge yourself in the documentation of the different classes PyNTA API.
If you want to start right away to improve the code, you can always look at the List of Todo’s.
Acknowledgements¶
This program was developed by Aquiles Carattino with the support of funding from NWO, The Netherlands Scientific Organization, under VICI grant (PI: Prof. Allard Mosk) and Projectruimte FOM.PR1.005 grant (PI: Dr. Sanli Faez) . This work was carried on at Utrecht University in the months between June 2018 and June 2019.